Genomic Study Reveals Unique Immune Signatures in Indigenous Indian Cattle
Research Summary: This study identifies breed-specific immune signatures in indicine breeds using integrative genomics, transcriptomics, and systems biology approaches, providing insights into biological significance underlying disease resistance and environmental adaptation.
Researcher Spotlight

Menaka Thambiraja is a researcher specializing in genetic variant discovery, multi-omics integration, and systems biology to explore complex biological mechanisms. She is passionate about integrating experimental and computational approaches to gain deeper insights into biological systems and is also interested in developing deep learning-based multimodal models to uncover meaningful patterns from biological data.
LinkedIn: www.linkedin.com/in/menakaraj128
Twitter: @MenakaRaj12
Instagram: rhia_aashi
Lab: Dr. Ragothaman M Yennamalli, SASTRA Deemed to be University
Lab website: https://yennamalli-lab.in/
What was the core problem you aimed to solve with this research?
Indicine cattle are well known for their natural disease resistance and ability to adapt to harsh environmental conditions, compared to crossbred and exotic breeds. In recent years, crossbreeding programs have primarily aimed to combine these resilience traits with improved milk production. While such programs have successfully enhanced milk yield, they have not consistently preserved the disease resistance and adaptability observed in pure indicine breeds. This limitation highlights the need to better understand the underlying genetic and genomic factors responsible for these important traits in indicine cattle.
In this context, our research focuses on unraveling the genetic architecture of immune-related traits in indicine breeds, as immune system function plays a central role in disease resistance. However, immune traits are highly complex, being influenced by multiple genes, regulatory factors, and evolutionary pressures. This complexity makes them difficult to study using traditional breeding methods or single-marker approaches. To address this challenge, we employed an integrative genome-wide comparative approach to identify key immune signatures and understand their functional significance in indicine breeds.
How did you go about solving this problem?
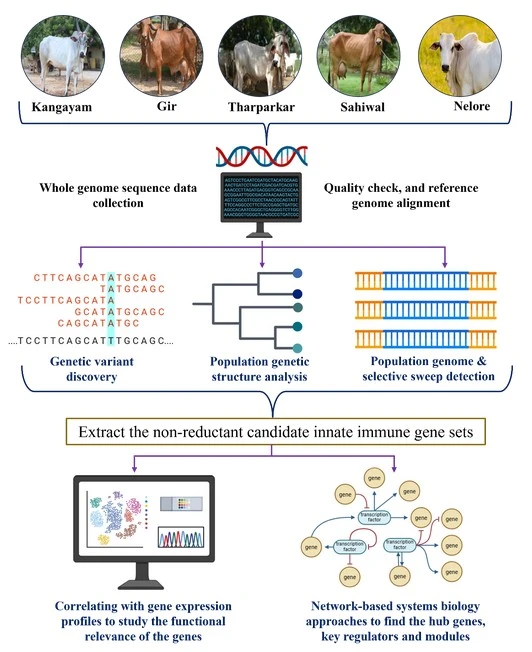
We began by collecting publicly available whole-genome sequencing data for several indicine cattle breeds, including Kangayam, Gir, Tharparkar, Sahiwal, and Nelore, from the NCBI Sequence Read Archive using SRA toolkit. Genomic variants within immune genes were identified using the Genomic Analysis Toolkit (GATK) and functional annotation were performed using SnpEff and SnpSift. We further applied population genomics approaches, including runs of homozygosity and selective sweep detection using RAiSD to identify the genomic regions shaped by immunity traits.
To expand our analysis, we performed whole genome sequencing of 108 animals representing four indicine breeds such as Kangayam, Gir, Tharparkar, and Sahiwal, along with a crossbred Karan Fries, and an exotic breed Holstein Friesian, using high-throughput sequencing platform as Illumina NovaSeq 6000. A comparative genome-wide analysis was then conducted across these groups using multiple variant callers, such as DELLY, FreeBayes, and GATK, to ensure robust identification of genomic variants. We also performed population genomics analysis such as genetic differentiation analysis (FST and dXY), and selective sweep detection, to identify the candidate innate immune genes. The candidate genes identified by genomic analysis were then correlated with gene expression profiles derived from peripheral blood mononuclear cells under both healthy and lipopolysaccharides-stimulated conditions, enabling us to understand their functional behavior during normal and immune-stimulated conditions. To further explore biological significance at systems level, we applied network-based systems biology approaches to identify key hub genes, central regulatory factors, and functional modules in indicine breeds. Finally, we developed a baseline deep-learning based multimodal model that integrates genomic variant and gene expression data to predict immune genes from non-immune genes, providing a foundation for identifying unique immunity-related patterns in indicine breeds.
“Our study discovers key immune signatures, hub genes, central regulators, and functional modules that could contribute to disease resistance and adaptation capability of indicine breeds” – Dr. Ragothaman M Yennamalli
How would you explain your research outcomes (Key findings) to the non-scientific community?
Our research identified a set of potential innate immune genes that show significant genomic differences in indicine breeds compared to crossbred and exotic breeds. Among them, innate immune genes such as CARD9 (rs132699263) and NLRP8 (rs42734179) carried disruptive variants in the whole-genome sequenced samples of all indicine breeds (Kangayam, Gir, Tharparkar, and Sahiwal), but were absent in crossbred (Karan Fries), and exotic breed (Holstein Friesian). These results were also consistent with the analysis of publicly available genomic data of Kangayam, Tharparkar, and Sahiwal that retrieved from NCBI, highlighting their reliability of carrying significant genomic differences. These innate immune genes play a key role in pathogen recognition receptor mediated signaling and innate immune activation.
We also found that genes involved in activating classical complement pathway and triggering immune defense against pathogens such as C1R, showed strong genetic divergence across all indicine breeds, indicating their significance in indicine-specific immunity. In addition, selective sweep detection revealed that genomic regions undergone natural selection and linked with adaptive immune traits, were identified in genes like C4BPA, TYRO3, and TP73, which are known to regulate immune responses and maintain immune homeostasis. These genes were uniquely observed in all indicine breeds, but were absent in crossbred.
To understand how the candidate innate immune genes identified from genome analysis could function in real biological conditions, we integrated gene expression data. We observed that innate immune gene LRP1B, which showed significant genomic differences, was also highly expressed in indicine breeds compared to crossbred under healthy conditions, highlighting its potential role in maintaining baseline immunity. Interestingly, this gene has also been previously reported as selection signatures in Vrindavani cattle, further highlighting its importance as a unique immunity-related signature in indicine breeds. In addition, analysis under immune-stimulated conditions using lipopolysaccharide treatment revealed that the gene LTB4R, having missense variant, was also consistently upregulated across all indicine breeds but not observed in crossbred, supporting their significance in indicine breed-specific immune activation. Furthermore, network-based systems biology approaches identified potential indicine breed-specific immune hub genes and central transcriptional regulatory factors that coordinate immune response. Among them, JAK2 identified as key hub gene present across all indicine breeds, while members of the STAT family were identified as central transcriptional regulators coordinating immune functions. Overall, our findings reveal a set of unique genetic and functional immune signatures that likely contribute to the disease resistance and environmental adaptability of indicine breeds.
What are the potential implications of your findings for the field and society?
Our findings identified a set of key innate immune genes in indicine breeds that provide a strong foundation for future functional validation and experimental studies. These genes can be further explored to better understand how genetic variations influence observable traits, particularly disease resistance and adaptation, through genotype-phenotype association studies.
Importantly, this knowledge can support the development of more informed and precise breeding strategies. By incorporating these immune-related genetic markers into selection programs, it may be possible to improve disease resistance and environmental adaptability in livestock, ultimately contributing to more sustainable and resilient livestock production systems.
What was the exciting moment during your research?
One of the most exciting moments in our study was identifying a set of innate immune genes that consistently showed significant genomic differences across all four indicine breeds such as Kangayam, Gir, Tharparkar, and Sahiwal. This consistency across multiple breeds strongly supported the presence of shared genetic mechanisms underlying immunity and disease resistance capability of indicine breeds.
Paper reference: Thambiraja M, Iyengar SK, Satishkumar B, Kavuru SR, Katari A, Singh D, Onteru SK and Yennamalli RM. (2026) Genetic basis of immunity in Indian cattle as revealed by comparative analysis of Bos genome. Scientific Reports 16, 11005. https://doi.org/10.1038/s41598-026-44002-9. https://www.nature.com/articles/s41598-026-44002-9.
Thambiraja M, Karthikeyan PP, Sezen M, Senthilkumar S, Singh D, Onteru SK, Yennamalli RM. (2025) ImmFinder: A Fully Connected Neural Network approach for predicting the immune genes in livestock. OMICS: A Journal of Integrative Biology 29(11):551-559. https://doi.org/10.1177/15578100251389910. https://journals.sagepub.com/doi/abs/10.1177/15578100251389910.
Thambiraja M, Roshan M., Onteru S.K, Yennamalli R.M. Integrative whole-genome analysis reveals genomic signatures of innate immunity in indicine cattle. biorxiv. https://doi.org/10.64898/2026.01.21.700751. Preprint – Under review in HLA: Immune Response Genetics journal.
Explore more
🎤 Career – Real career stories and job profiles of life science professionals. Discover current opportunities for students and researchers.
💼 Jobs – The latest job openings and internship alerts across academia and industry.
🛠️ Services – Regulatory support, patent filing assistance, and career consulting services.