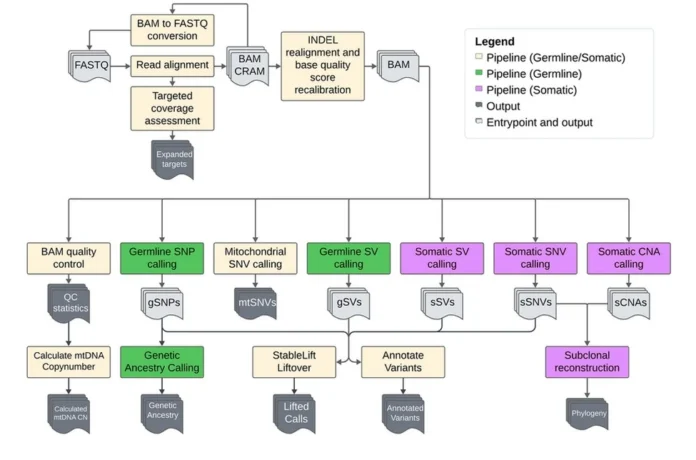
Metapipeline-DNA Automates Genomics Workflows for Scalable DNA Analysis
Research Summary: Metapipeline-DNA is an automated, configurable pipeline that transforms raw DNA sequencing data into comprehensive genetic and evolutionary insights across germline and tumor analyses.
Researcher Spotlight

Yash Patel is a cloud and AI infrastructure architect working on computational genomics research. He specializes in scalable bioinformatics pipelines, analysis automation, infrastructure optimization, and reproducible, high-throughput genomics workflows.
Linkedin: https://www.linkedin.com/in/yash-patel-52128b160
Lab: Prof. Paul Boutros, Sanford Burnham Prebys Medical Discovery Institute
Lab social media: https://x.com/theboutroslab
What was the core problem you aimed to solve with this research?
With the growing complexity of technologies and data volume and lowering costs of sequencing, DNA sequencing analyses require complex workflows involving multiple tools. This leads to a need to produce standardized, scalable, and robust workflows that can handle the large volume of generated data in a reproducible manner while scaling across the different compute environments available today.
How did you go about solving this problem?
We took the approach of architecting and developing an extensible Nextflow-based pipeline that integrates multiple algorithms into modular workflows. We placed a firm emphasis on making the pipeline scalable for processing large cohorts and easily extensible to maintain upkeep with novel algorithms as they become available. We automated the entire process from raw read alignment to variant calling, quality control, and visualization to allow processing of entire cohorts without input from users throughout the pipeline run. The design of metapipeline-DNA focused on enabling flexibility and robustness to different compute environments on which the pipeline may be run.
The goal is to automate quality control, determination of genetic variants and all the other analysis steps to make it much easier so that researchers do not need to write their own code to process their data. – Prof. Paul Boutros
How would you explain your research outcomes (Key findings) to the non-scientific community?
Metapipeline-DNA automatically processes DNA data to reliably and reproducibly identify genetic changes and tumor evolution patterns. These features guide research into identifying the causes of cancer and how it evolves to ultimately develop effective therapies for cancer.
What are the potential implications of your findings for the field and society?
The finding and work lower the barrier to performing complex genomic analyses by unifying different algorithms into a single, automated workflow. It improves reproducibility and consistency across studies. By enabling scalable, cloud-enabled analysis, it supports population-level genomics and precision medicine efforts that rely on large cohorts of data. It also allows researchers to rapidly integrate novel algorithms modularly to accelerate genomics and cancer research.
What was the exciting moment during your research?
An exciting moment was seeing the results of implementing consensus-based variant calling across different types of variants and seeing the reduction in false positives without sacrificing true discoveries. It demonstrated the utility of integrating multiple algorithms to produce more reliable results.
Paper reference/citation: Patel, Y., Zhu, C., Yamaguchi, T. N., et al. (2026). Metapipeline-DNA: A comprehensive germline and somatic genomics Nextflow pipeline. Cell Reports Methods, 101340. https://doi.org/10.1016/j.crmeth.2026.101340
Explore more
🎤 Career – Real career stories and job profiles of life science professionals. Discover current opportunities for students and researchers.
💼 Jobs – The latest job openings and internship alerts across academia and industry.
🛠️ Services – Regulatory support, patent filing assistance, and career consulting services.