HER2 Mutations Drive Drug Resistance in Breast Cancer, Study Reveals New Treatment Strategies
Research Summary: HER2 dimerization-domain mutations drive receptor switching from HER2 homodimers to HER2–HER3 heterodimers, activating AKT signaling and conferring resistance to trastuzumab and neratinib, while retaining sensitivity to tucatinib and ADCs.
Researcher Spotlight

Pranay Dey was a former doctoral student from the Molecular Functional Imaging Lab aka De lab at ACTREC, TMH, and is currently a postdoctoral fellow at EPFL, Switzerland, studying the role of androgen receptor signaling in early breast carcinogenesis.
Linkedin: https://www.linkedin.com/in/pranay-dey-ph-d-16b196280/
Instagram: pranay_dey_94
Lab: Dr. Abhijit De, ACTREC, TMH, HBNI
Lab social media: https://www.linkedin.com/in/abhijit-de-301827b/
https://x.com/DeLab5038 | https://x.com/Abhijit75960170
Lab website: https://www.mfil-actrec.com/
What was the core problem you aimed to solve with this research?
The core problem we aimed to address was the persistent challenge of therapeutic resistance in HER2-positive breast cancer, where many patients initially respond to targeted therapies but relapse within a short period. Despite the clinical success of drugs like trastuzumab, the mechanisms driving this resistance remain incompletely understood, limiting our ability to design more effective and durable treatments. In this study, we focused on an underexplored aspect of HER2 biology i.e. mutations in its dimerization domains, which are critical for how the receptor interacts with other partners on the cell surface. We sought to determine whether these mutations could fundamentally alter receptor behavior, rewire downstream signaling pathways, and enable cancer cells to bypass therapeutic inhibition. In a broader sense, our goal was to understand how subtle structural changes in a key oncogenic receptor can reshape cellular communication networks, drive aggressive tumor behavior, and compromise treatment efficacy. By uncovering these mechanisms, we aimed to provide a foundation for more precise therapeutic strategies that can overcome resistance and improve outcomes for patients with HER2-positive breast cancer
How did you go about solving this problem?
To address this problem, we took a multidisciplinary approach combining patient data analysis, modeling of HER2 and HER3 protein structures, and experimental validation. We began by mining large-scale breast cancer datasets (TCGA and METABRIC) to identify recurrent mutations in the HER2 dimerization domains and assess their clinical relevance, including their association with patient outcomes. We then used molecular dynamics simulations and structural modeling to understand how these mutations alter HER2 structure and its homo-dimer stability. This enabled us to generate a mechanistic hypothesis that these mutations might disrupt HER2 homodimer formation and instead promote HER2-HER3 heterodimerization. To validate this, we engineered breast cancer cell models expressing either wild-type HER2 or specific mutants. Using proximity ligation assays and co-immunoprecipitation, we demonstrated a clear shift in receptor interactions. We further characterized downstream signaling shift using phospho-protein analyses and live-cell assays, revealing a switch from ERK to AKT signaling.
Finally, we evaluated the functional and therapeutic consequences of these mutations using invasion assays and in vivo tumor models. Importantly, we tested clinically relevant therapies and found that while these mutations confer resistance to trastuzumab and neratinib, the cells remain sensitive to tucatinib and antibody-drug conjugates (ADCs) like TDxd and TDM1, highlighting alternative therapeutic strategies for patients suffering from drug resistance.
“HER2 therapies improved outcomes, but this study shows single mutations can rewire tumor biology, highlighting need for patient-specific treatment strategies.” – Dr. Abhijit De
How would you explain your research outcomes (Key findings) to the non-scientific community?
Our research addresses why some breast cancer treatments stop working after an initial response. We focused on a receptor protein called HER2, which is a key driver in a major subset of breast cancer cases and is commonly targeted by existing therapies. We found that small mutations in HER2 can change how it interacts with other proteins on the cell surface. Instead of functioning in the usual way, the protein essentially “switches partners” and activates an alternative survival pathway. This allows cancer cells to bypass the effects of standard treatments and continue growing. What’s important is that this gives us a clearer explanation of how resistance develops at a very fundamental level. Even more encouraging, we showed that although these mutations make some commonly used drugs less effective, newer therapies such as tucatinib and certain antibody-based treatments can still target these cancer cells successfully.
So overall, our work not only explains why treatment resistance occurs, but also helps guide more effective treatment choices, with the potential to improve outcomes for patients.
What are the potential implications of your findings for the field and society?
Our findings have important implications for both the scientific field and patient care. At a fundamental level, we show that resistance in HER2-positive breast cancer can arise from structural changes in the HER2 receptor itself, which alter how it interacts with other proteins and rewire downstream signaling. This adds a new layer to our understanding of resistance, moving beyond compensatory pathway activation to the level of receptor dynamics and interaction networks.
From a clinical perspective, this highlights the importance of profiling these specific HER2 mutations in patients before prescribing medications. In simpler terms, it means that not all HER2-positive cancers behave the same way, some may “adapt” and find alternative survival routes, making certain treatments less effective. Now, by recognizing the presence of these mutations early could help doctors choose therapies that are more likely to work, such as tucatinib or antibody-drug conjugates, rather than relying on a trial-and-error approach, commonly used in clinics.
More broadly, our work strongly supports the shift toward precision medicine where treatment is tailored based on the molecular features of each patient’s tumor. By better understanding how and why resistance occurs, we can improve treatment selection, reduce ineffective therapies, and ultimately enhance outcomes and quality of life for patients.
What was the exciting moment during your research?
There were several exciting moments throughout the project, but if I had to highlight three, they would be key points where our hypotheses were clearly validated. The first was when I performed the initial co-immunoprecipitation experiment. It was particularly exciting because it provided the first experimental evidence supporting our simulation data, confirming that the receptor interactions were indeed changing as we had predicted from our in-silico work. The second was when I performed the BRET assay, which allowed me to observe downstream signaling in real time in live cells. Seeing a clear switch from ERK to AKT signaling was a powerful moment, as it directly connected the structural changes to functional consequences. Finally, the proximity ligation assay (PLA) was especially rewarding. Not only did it produce visually striking results, but it also added a strong spatial and quantitative layer of validation to our findings, reinforcing the overall story.
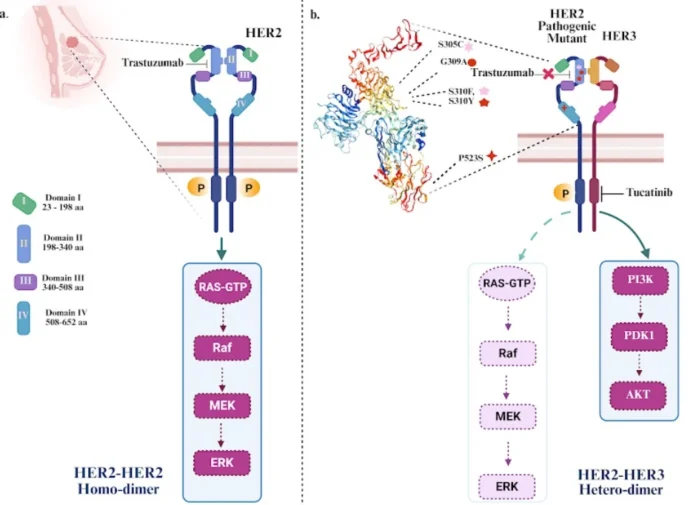
Figure Caption: Schematic explaining change in HER2 dimerization status and downstream signaling switching due to presence of specific mutations in the extracellular domains of the receptor. a In HER2-positive breast cancer with overexpression of the wild-type HER2 receptor, HER2:HER2 homo-dimer forms leading to constitutive activation of downstream ERK signaling. The monoclonal antibody drug, trastuzumab, binds to the extracellular domain of HER2 can prevent the dimerization and therefore downstream signaling activation. b In presence of hotspot mutations in the dimerization domain II and IV (i.e., G309A, S310F, S310Y, P523S), structural change in the dimer face of HER2 potentiate HER2:HER3 heterodimerization and activate both PI3K–AKT signaling axis, thereby turn trastuzumab treatment ineffective. In the altered scenario, as the small molecule tucatinib inhibits kinase activity of HER2:HER3 heterodimer, this drug shows growth inhibitory effect.
Paper reference: Dey, P., Bishnu, A., Patel, J. et al. Hotspot mutations in HER2 interfaces destabilize structure, causing breast cancer treatment failure. Oncogene 45, 414–430 (2026). https://doi.org/10.1038/s41388-025-03653-0.
Explore more
🎤 Career – Real career stories and job profiles of life science professionals. Discover current opportunities for students and researchers.
💼 Jobs – The latest job openings and internship alerts across academia and industry.
🛠️ Services – Regulatory support, patent filing assistance, and career consulting services.