New Drug-Resistant Bacterium Discovered in Silkworms Raises Public Health Concerns
Research Summary: This study reports a novel extensively drug-resistant bacterium, Stenotrophomonas raiganjensis, from silkworm hemolymph, characterized using genomic and polyphasic taxonomy under SeqCode, highlighting its uniqueness, resistance traits, and clinical significance.
Researcher Spotlight

Dr. Rittick Mondal completed his PhD at the Department of Sericulture, Raiganj University, under Dr. Amit Kumar Mandal, and is currently an Assistant Professor at IQ City Medical College Hospital.
Lab: Dr. Amit Kumar Mandal, Raiganj University, West Bengal
What was the core problem you aimed to solve with this research?
Sericulture is a cottage-based industry that sustains the livelihood of many rural families, where even small disease outbreaks in silkworms can lead to significant economic losses. However, the exact microbial agents responsible for these infections are often not well characterized, especially in local rearing environments. The core problem we aimed to address was to identify and characterize the microorganisms associated with diseased silkworms and understand their biological and clinical significance.
In particular, we were interested in determining whether these pathogens possess antimicrobial resistance traits, which could make them difficult to control and potentially hazardous beyond the sericulture system. Another important concern was whether such bacteria could pose a risk to humans who are in close contact with infected larvae during rearing and handling. Therefore, our study also indirectly highlights the need for awareness, proper hygiene practices, and biosafety measures while handling diseased silkworms, especially when dealing with drug-resistant microorganisms.
Overall, the research bridges gaps between sericulture health management, rural economic stability, and emerging public health concerns related to antimicrobial resistance.
“Our study demonstrates how genome-based taxonomy can uncover novel drug-resistant bacteria from overlooked niches, linking environmental microbiology with emerging public health concerns.” – Dr. Amit Kumar Mandal
How did you go about solving this problem?
To address this problem, we collected diseased silkworm larvae from the silkworm rearing house of Raiganj University and isolated bacteria from their hemolymph under sterile conditions. The isolated strain was then studied using a comprehensive polyphasic approach, integrating classical microbiology with advanced molecular and genomic techniques.
We performed morphological, biochemical, and physiological characterization, along with antimicrobial susceptibility testing to determine its resistance profile. Further, we carried out 16S rRNA gene sequencing, multilocus sequence analysis (MLSA), and whole-genome sequencing. Genome-based metrics such as average nucleotide identity (ANI) and digital DNA–DNA hybridization (dDDH) were used to confirm its novelty. Additionally, we analyzed the genome to identify antimicrobial resistance and virulence-associated genes.
By combining these approaches under the SeqCode framework, we were able to accurately identify, classify, and assess the potential risks of this novel drug-resistant bacterium.
How would you explain your research outcomes (Key findings) to the non-scientific community?
In simple terms, we discovered a new type of bacteria living inside diseased silkworms that is resistant to many commonly used antibiotics. This means it is harder to control and could spread if not handled carefully. Using advanced DNA-based techniques, we confirmed that this bacterium is completely new and different from known species.
Our findings suggest that organisms present in silkworm rearing environments are not only important for crop health but could also have implications for human health, especially for people who are in close contact with infected larvae. This highlights the importance of proper hygiene, awareness, and safe handling practices in sericulture to prevent potential risks.
What are the potential implications of your findings for the field and society?
Our findings have important implications for both science and society. For the scientific community, this study contributes to microbial taxonomy by introducing a novel bacterial species using modern genome-based classification. It also adds valuable insights into antimicrobial resistance, especially in lesser-explored environments like insect systems.
For society, particularly in sericulture-dependent rural areas, the discovery highlights the need for better disease monitoring and management practices to reduce economic losses. The presence of an extensively drug-resistant bacterium also raises awareness about potential health risks for people handling infected silkworms. This emphasizes the importance of adopting proper hygiene, biosafety measures, and responsible antibiotic use to prevent the spread of resistant microorganisms.
What was the exciting moment during your research?
The most exciting moment came when the genome analysis results clearly showed that our isolate did not match any known species. What initially seemed like a routine bacterial isolation from infected silkworms turned into the discovery of a completely novel organism.
It became even more special when we confirmed that this new bacterium was extensively drug-resistant, highlighting its potential significance. Adding to that excitement, we had the opportunity to honor Raiganj town by naming the species Stenotrophomonas raiganjensis, making the discovery personally meaningful as well as scientifically important.
That moment-combining novelty, impact, and a connection to our place of work—was truly unforgettable.
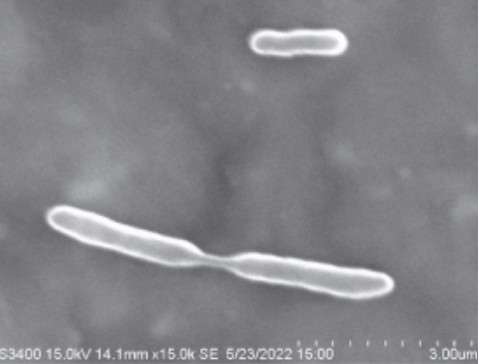
Figure Caption: Scanning Electron Micrograph of Stenotrophomonas raiganjensis at 15,000 × magnification
Paper reference: Mondal, R., Manna, A., Mandal, P., Kurt, H., Sen, A., Jangid, K., & Mandal, A. K. (2026). Stenotrophomonas raiganjensis sp. nov., an extensively drug-resistant bacterium isolated from Bombyx mori L., described under the SeqCode. Systematic and Applied Microbiology, 126699. https://doi.org/10.1016/j.syapm.2026.126699
Explore more
🎤 Career – Real career stories and job profiles of life science professionals. Discover current opportunities for students and researchers.
💼 Jobs – The latest job openings and internship alerts across academia and industry.
🛠️ Services – Regulatory support, patent filing assistance, and career consulting services.