Renoir, A New Computational Tool Developed at IIT Kanpur Charts Cell-to-Cell Communication Inside a Tissue
Researchers from Indian Institute of Technology Kanpur and the Garvan Institute of Medical Research develop Renoir, an AI-driven framework that maps cell–cell signalling across the topography of a tissue — revealing new insights in fetal liver development and liver cancer.
Cells inside tissues are constantly exchanging chemical messages that control development, immunity, and disease. But understanding exactly which cells are communicating — and how those signals influence neighbouring cells — has remained a major challenge in biology. Now, researchers from the Indian Institute of Technology Kanpur and the Garvan Institute of Medical Research have developed Renoir, a computational framework that maps these cellular conversations across the spatial landscape of tissues. The study, led by Hamim Zafar (IIT Kanpur), could advance understanding of fetal liver development while also revealing new therapeutic opportunities in cancer biology.
Every tissue in the body is a bustling community where cells constantly exchange signals to coordinate essential processes such as tissue development and immune responses. When these conversations go awry, diseases like cancer can develop and spread. Modern technologies such as spatial transcriptomics now allow scientists to measure gene activity in thousands of cells while preserving their precise location within a tissue — producing a high-resolution map of a tumour, brain region, or a developing organ. However, it has remained difficult to understand how signals from one cell actually influence the behavior of the recipient cells, which downstream genes are switched on by the signals, especially within the complex spatial organization of tissues.
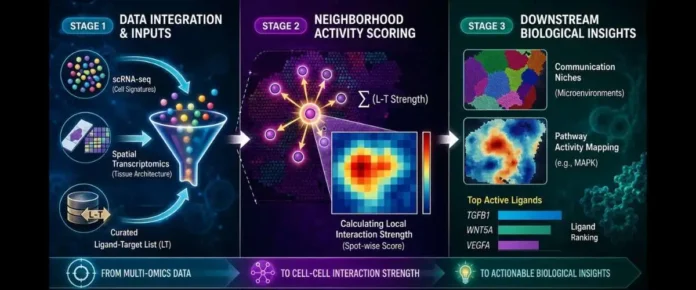
Named after the painter who captured intricate social scenes, Renoir (short for ligand-taRget intEractions across spatial topogRaphy) assigns a “neighbourhood activity score” to every plausible ligand–target gene pair at every location in a tissue. Renoir calculates a “neighbourhood activity score” for signalling interactions across tissues. The framework considers which cell types are present, whether nearby cells carry the appropriate receptors to receive a signal, and how ligand–target relationships appear in single-cell datasets. Renoir also enables the identification of “communication niches” in the tissue, where particular signaling pathways are active.
Checking for the presence of the appropriate receptor on the receiving cell is what sets Renoir apart. This simple but crucial safeguard prevents the tool from flagging communications that physiologically cannot occur — a major pitfall of earlier methods. When tested across more than 200 simulated datasets spanning brain, breast cancer, and intestinal tissues, Renoir outperformed leading tools such as SpatialDM, COMMOT, and stLearn, achieving up to a 98% improvement in spatial accuracy.
The team applied Renoir to four very different biological systems, each highlighting a unique strength of the framework. Renoir delineated 20 distinct communication niches aligned with known anatomical regions of the brain and showed how specialised astrocytes in the cortex, hippocampus, thalamus, and white matter converse with neighbouring neurons and oligodendrocytes — producing a region-by-region atlas of neuro-glial communication. In one of the most aggressive forms of breast cancer, Renoir uncovered sub-neighbourhoods within tumours where cancer cells, immune cells, and connective tissue cells exchange distinct sets of chemical messages. Some pockets harboured signals known to drive tumour spread (such as TGFB3 and CCL2), while others showed immune-suppressive dialogues (IL-6/JAK/STAT3 signalling). Such fine-grained maps could help researchers design therapies that target the specific interaction in specific regions of the tumour.
The IITK team collaborated with Prof. Ankur Sharma’s laboratory at the Garvan Institute of Medical Research, Australia, where spatial transcriptomic datasets from fetal liver and liver cancer were generated to evaluate Renoir. Using the cutting-edge 10x Visium HD platform, Renoir revealed a previously unrecognized interaction between liver cells (hepatocytes) and resident embryonic macrophages, mediated by a protein called PLG that is important for liver regeneration. The team showed that a specific subset of PLG-positive hepatocytes co-localises with FOLR2-positive macrophages, suggesting a specialised niche during organ development.
The most striking finding came from liver cancer. Applying Renoir to single-cell-resolution spatial data from hepatocellular carcinoma patients, the researchers predicted that onco-fetal cells in the tumour microenvironment reprogram a stem-like cancer cell population (called MUC6-positive bipotent cells). These onco-fetal cells include fetal-like fibroblasts, macrophages, and endothelial cells. The team then put the prediction to the test in the laboratory: adding IL6 to liver cancer cells caused a dose-dependent expansion of CD133-positive stem-like cells — the very population thought to drive tumour relapse and resistance to therapy.
Renoir works across the major spatial transcriptomics platforms used by laboratories worldwide — including 10x Visium, Visium HD, NanoString CosMx, and 10x Xenium — and is freely available as open-source software on GitHub. By revealing which cell-to-cell signals are active in which regions of a tissue, Renoir could accelerate discoveries in cancer biology, developmental research, and precision medicine. As spatial transcriptomics technologies become increasingly widespread, frameworks like Renoir could help researchers move beyond descriptive maps of tissues toward a mechanistic understanding of how cells coordinate health and disease.
Reference study: Rao N, Kumar T, Kazemi D, et al. Charting spatial ligand-target activity using Renoir. Nature Communications (2026). DOI: 10.1038/s41467-026-72388-7
Code availability: github.com/Zafar-Lab/Renoir (MIT licence)
New Book Launched – Molecules, Mentors & Mindsets: Building Indian Biopharma
Buy your copy today: https://biopatrika.com/science-society/book-molecules-mentors-mindsets-building-indian-biopharma-biocon/
Book Launch: Molecules, Mentors & Mindsets: Building Indian Biopharma | Biocon Focus